This post is Part 2 in a series on how to configure a system for LLM deployments and development usage. Part 2 is about installing and configuring container tools, Docker and NVIDIA Enroot.


This post is Part 2 in a series on how to configure a system for LLM deployments and development usage. Part 2 is about installing and configuring container tools, Docker and NVIDIA Enroot.

This post is Part 1 in a series on how to configure a system for LLM deployments and development usage. The configuration will be suitable for multi-user deployments and also useful for smaller development systems. Part 1 is about the base Linux server setup.

This is a short note on setting up the Apache web server to allow system users to create personal websites and web apps in their home directories.
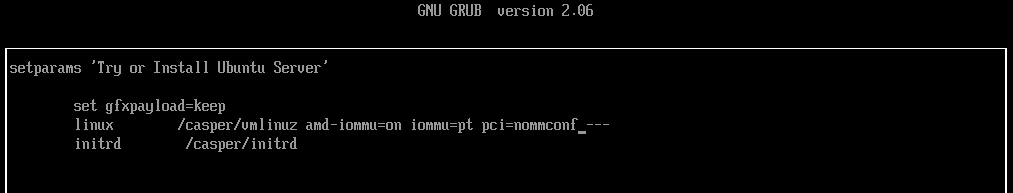
This post is a short HowTo on passing Linux kernel boot options during OS installation and persisting them for future system starts

Learning go (Golang) is one of my resolutions for 2023. It looks like a great cross platform compiled language with a straightforward simple syntax with modern features. I have multi-OS projects in mind where I expect it to be ideal. So, I’ll get started …
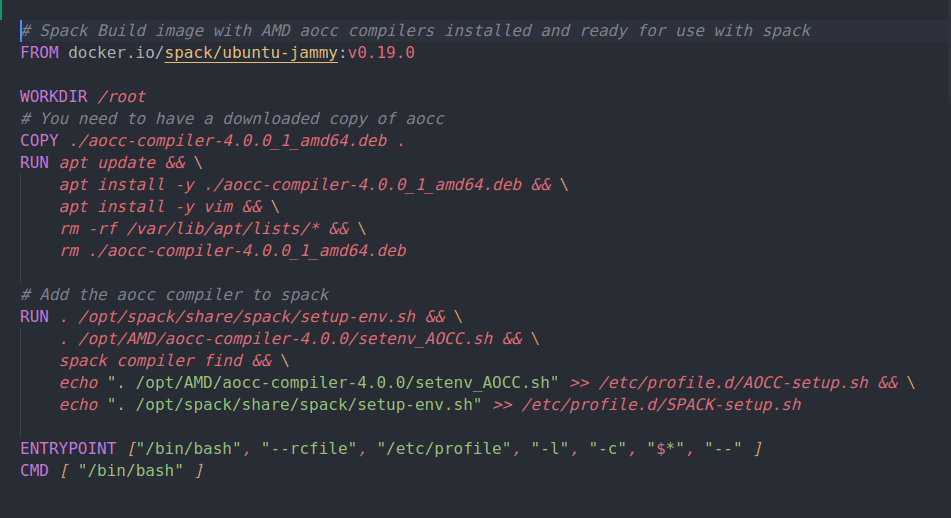
AMD has recently released version 4.0 of their AOCC compiler which includes support for AVX512 on the Zen4 architecture. This post details building a Docker image containing the Spack package manager/build system together with AMD AOCCv4.0.0 compilers. This will be used as the build image for multi-stage Dockerfiles that will be used to compile scientific applications and benchmarks with targeted Zen3/4 optimizations. It is the first step in that process.
This post is a follow up to How-To: Make Ubuntu Autoinstall ISO with Cloud-init written in Sept. 2021. We will look at changes needed for Ubuntu 22.04.
This is just a short post to announce a more usable version of the NVIDIA GPU powerlimit setup script that I released a few months ago. This update to version 0.2 uses an interactive mode to set GPU powerlimits and optionally setup a systemd unit file to set these limits on subsequent reboots.
In this post we look at using a testing Lab of Windows systems as a benchmarking platform for Linux scientific application using network boot with nfsroot and home mounts. Linux is boot on the systems “diskless” leaving the Windows installs untouched. LTSP turned out to be a great time saver for setting up the configuration.
This post presents testing data showing that power-limit reduction on NVIDIA GPUs have give significant benefits for both high wattage and lower wattage GPUs. Power-limit vs Performance data is presented for 1-4 A5000 and 1-4 RTX3090 GPUs.