GTC was intense, and there is a massive debate raging right now: Is AI a bubble? Is Jensen just a shovel salesman? Or are we looking at the new bedrock?


GTC was intense, and there is a massive debate raging right now: Is AI a bubble? Is Jensen just a shovel salesman? Or are we looking at the new bedrock?

This guide will walk you step-by-step through installing and configuring our tailored AI environments on your Puget Systems workstation or server.

We look at AI the way our customers do, which is why we built the Puget Systems Docker App Packs: to help you get up and running with AI inference fast!

How I used “Vibe Coding” and 25 years of experience to tame a liquid-cooled supercomputer in two weeks.

A brief look into using a hybrid GPU/VRAM + CPU/RAM approach to LLM inference with the KTransformers inference library.
An introduction to NPU hardware and its growing presence outside of mobile computing devices.

Presenting local AI-powered software options for tasks such as image & text generation, automatic speech recognition, and frame interpolation.
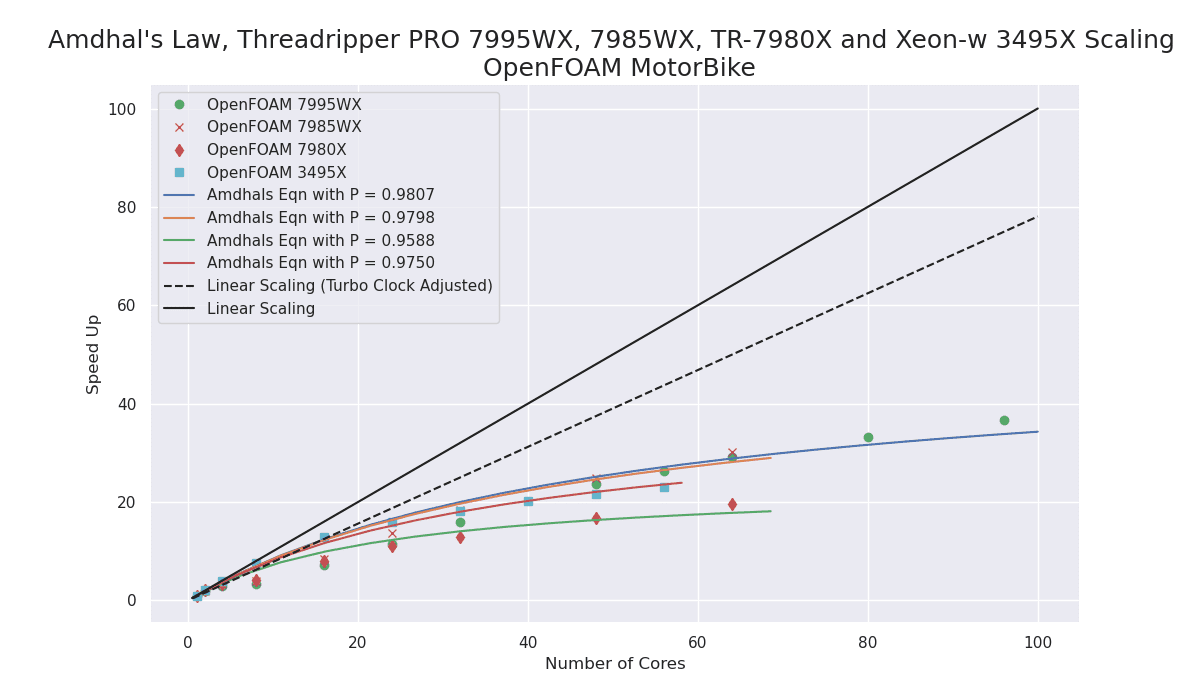
The performance improvement with the new Zen4 TrPRO over the Zen3 TrPRO is very impressive!
My first recommendation for a Scientific and Engineering workstation CPU would now be the AMD Zen4 architecture as either Zen4 Threadripper PRO or Zen4 EPYC for multi-socket systems.

Evaluating the speed of GeForce RTX 40-Series GPUs using NVIDIA’s TensorRT-LLM tool for benchmarking GPU inference performance.

Results and thoughts with regard to testing a variety of Stable Diffusion training methods using multiple GPUs.